DKFZ/NCT/DKTK MASTER Program: Handbook on the use of genomic data in precision oncology
IIn precision oncology, often referred to as personalized cancer medicine, the analysis of selected cancer genes provides important information that can be used to plan the most targeted treatment possible. However, such analysis does not capture all molecular characteristics and individual vulnerabilities of tumors. This is where comprehensive molecular characterization can add critical clinical value for diagnostics, disease progression prediction, and treatment decisions. However, the complexity and heterogeneity of the resulting data is challenging, and to date, no general standards exist for the use of genomic data in precision oncology. The recent publication by the team of the DKFZ/NCT/DKTK MASTER program (Molecularly Aided Stratification for Tumor Eradication Research) aims to fill this gap.
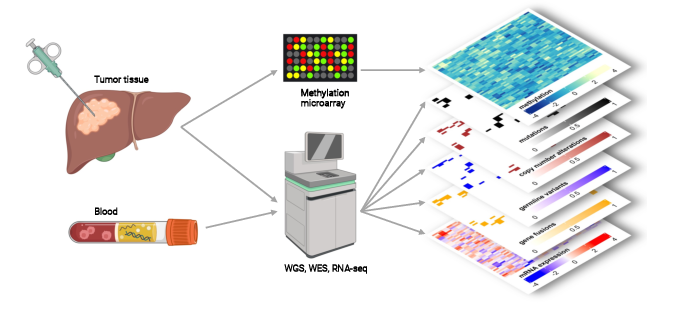
The "Handbook for the Interpretation of Whole Genome, Transcriptome and Methylome Data for Precision Oncology" shows how scientists at the NCT Heidelberg and NCT Dresden deal with very comprehensive molecular data in clinical applications within the framework of the MASTER program. The entire workflow is illustrated, from the collection of tumor tissue, molecular and bioinformatic analysis and prioritization of molecular alterations to the assignment of treatment recommendations and discussion in the molecular tumor board. Case studies provide insight into the approach to multidimensional tumor profiles and guidance on the interpretation of their biological impact and diagnostic and therapeutic implications.
This type of diagnostic approach is unique in Germany to date. The handbook is intended to support physicians, biologists and bioinformaticians in the clinical interpretation of genome-wide molecular information.
It is published in the journal NPJ Precision Oncology and available here
More about the DKFZ/NCT/DKTK-MASTER program can be found here







