DISSECT
Tumor Cell Determinants of Oncolytic Immunotherapy
PI(s)
Christine E. Engeland
Contributing CCU members
Sophie C. S. Pernickel, Nicolas Duus, Theresa E. Schäfer, Jessica Albert, Birgit Hoyler, Stefanie Sawall
Cooperation partners
Dr. Claudia Ball, National Center for Tumor Diseases Dresden and German Cancer Research Center Heidelberg
Priya Chudasama and Stefan Fröhling, German Cancer Research Center
Jessica Hassel, National Center for Tumor Diseases and University Hospital Heidelberg
Benedikt Brors, Sebastian Uhrig, and Dr. Jennifer Hüllein, German Cancer Research Center Heidelberg
Anja Ehrhardt and Wenli Zhang, Witten/Herdecke University
Anna Hartley and Antonio Marchini, German Cancer Research Center Heidelberg
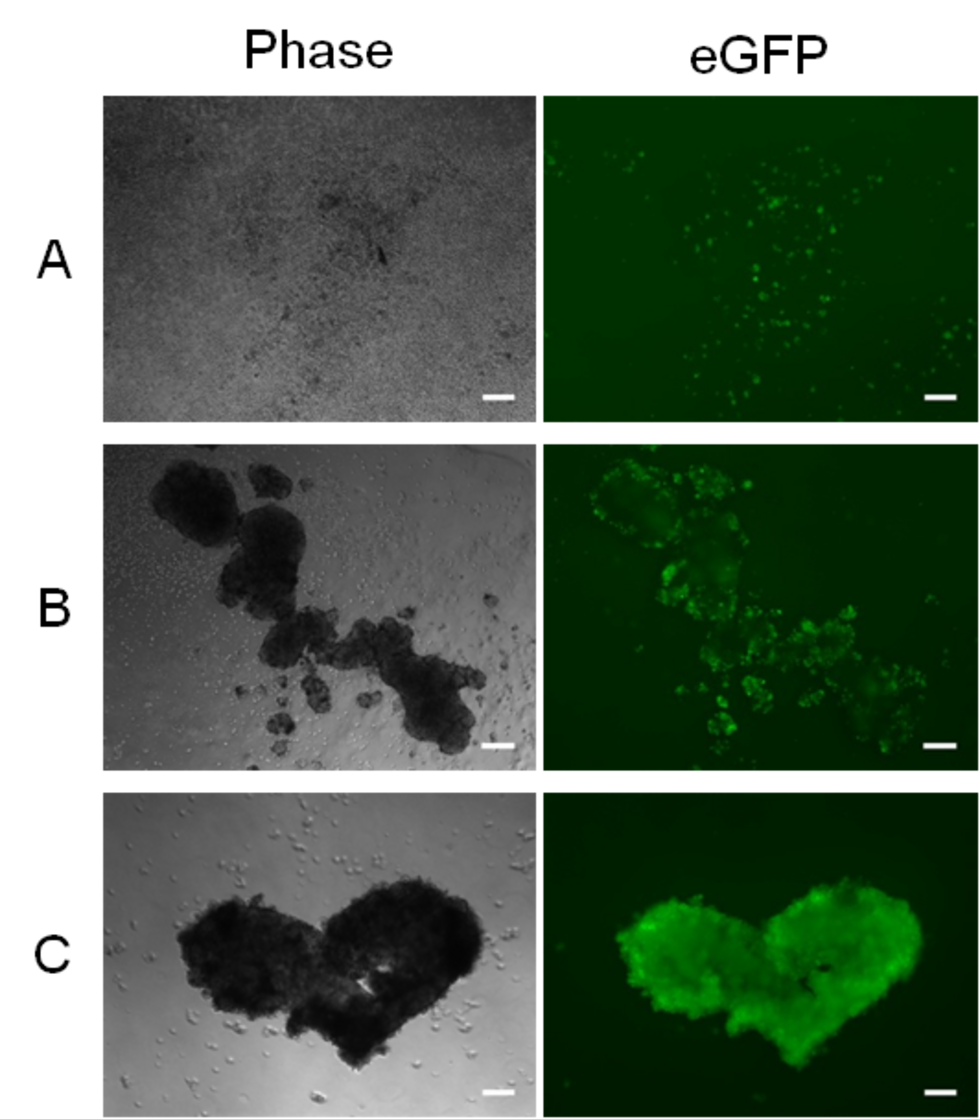
Several oncolytic viruses are currently advancing in clinical development. However, biomarkers to predict which patients will benefit most from these novel therapeutics are currently lacking. We study patient-derived models to identify tumor-intrinsic factors associated with response or resistance to oncolytic virotherapy with a focus on, but not limited to, the measles vaccine strain.
Collaboration partners in clinical and translational research include the group of Dr. Claudia Ball (NCT Dresden), Dr. Priya Chudasama (Precision Sarcoma Research, DKFZ and NCT Heidelberg), Prof. Dr. Stefan Fröhling (Molecular and Cellular Oncology, DKFZ and NCT Heidelberg), and Prof. Dr. Jessica Hassel (Dermato-Oncology, NCT Heidelberg). Bioinformatic experts working on this project are Dr. Mario Huerta (Translational Medical Oncology, NCT Heidelberg), Prof. Dr. Benedikt Brors, Sebastian Uhrig, and Dr. Jennifer Hüllein (Applied Bioinformatics, DKFZ).
Publications
- Mechanisms of measles virus oncolytic immunotherapy. Pidelaserra-Martí G, Engeland CE. Cytokine & Growth Factor Reviews 2020 Jul 3:S1359-6101(20)30175-1.
- Targeted BiTE Expression by an Oncolytic Vector Augments Therapeutic Efficacy Against Solid Tumors. Speck T, Heidbuechel JPW, Veinalde R, Jaeger D, von Kalle C, Ball CR, Ungerechts G, Engeland CE. Clinical Cancer Research 2018; 24: 2128-37.
- CTLA-4 and PD-L1 checkpoint blockade enhances oncolytic measles virus therapy. Engeland CE, Grossardt C, Veinalde R, Bossow S, Lutz D, Kaufmann JK, Shevchenko I, Umansky V, Nettelbeck DM, Weichert W, Jäger D, von Kalle C, Ungerechts G. Molecular Therapy 2014 Nov;22(11):1949-59.
Funding
Deutsche Krebshilfe, Mildred Scheel MD Fellowships (to ND and TES)
Systems Immunodiagnostics Platforms for Adaptive Clinical Trials and Personalized Immuno-oncology
PI(s)
Thomas Walle, Guy Ungerechts
Contributing CCU members
Sunanjay Bajaj, Birgit Hoyler, Stefanie Sawall
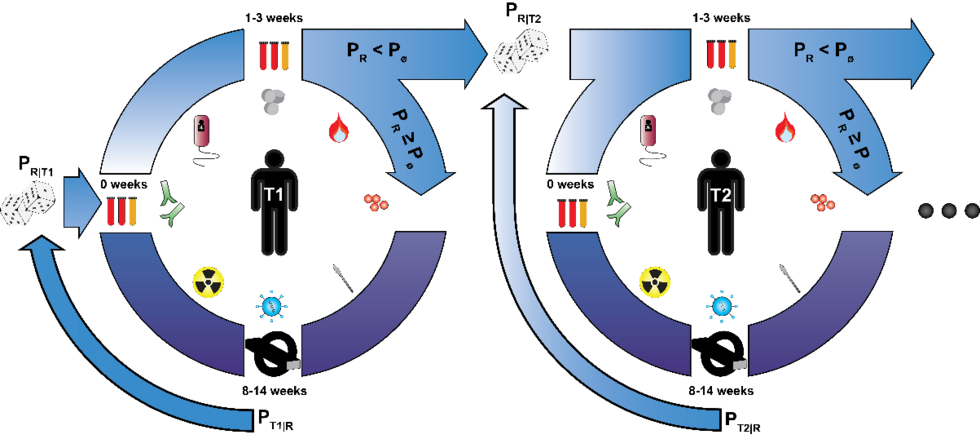
Combination cancer immunotherapies hold promise to dramatically improve overall survival of cancer patients and can readily be encoded in viral vectors. However, the number of possible combinations exceeds the numbers of feasible clinical trials. Even adaptive clinical trials which rapidly modify the trial protocol based on patients' outcomes fail to deliver the speed required to screen enough combination therapies.
We and others have shown that similar immune responses can be observed across multiple types of tumor and combination immunotherapies. Here we will harness these patterns to create a pan-solid tumor, low-cost, multiparametric and interpretable cellular biomarker model in cancer patients across multiple combination cancer immunotherapies in the NCT ANTICIPATE trial.
This biomarker model will allow for fast response assessment and rapid cycling of cancer patients through multiple therapies until an effective therapy is found. For this purpose, we will analyze changes in peripheral blood mononuclear cell (PBMC) phenotype under steady state and under controlled ex vivo perturbation using CITE-seq (10.1038/nmeth.4380) and flow cytometry with rational panels driven by feature selection methods. We will also analyze the serum proteome PBMCs are embedded in by mass spectrometry diligently working together with the laboratory of Prof. Dr. Jeroen Krijgsveld at DKFZ. In collaboration with the laboratory of Dr. Dana Pe'er at the Memorial Sloan Kettering Cancer Center (New York, U.S.A.) we will use these omics data to construct a machine learning classifier to accelerate cancer immunotherapy development in adaptive clinical trials by a manifold and head towards personalized virotherapy immunomodulation.
Publications
- Cytokine release syndrome-like serum responses after COVID-19 vaccination are frequent and clinically inapparent under cancer immunotherapy. Walle T, Bajaj S, […], Hoyler B, Zielske L, Skatula R, Sawall S, Leber MF, […], Halama N, Springfeld C, Kirsten R, Hassel JC, Jäger D, NCT ANTICIPATE Investigators & Ungerechts G. Nature Cancer. 2022
- Immune profiling of human tumors identifies CD73 as a combinatorial target in glioblastoma. Goswami S*, Walle T*, Cornish AE*, Basu S*, [...], Sharma P. Nature Medicine. 2020; 26: 39-46. *equal contribution
Funding
DKFZ Clinician Scientist Program (Dieter Morszeck Stiftung, to TW)







